Lomb-Scargle Periodograms for Multiband Data#
The Lomb-Scargle periodogram (after Lomb [1], and Scargle [2]) is a commonly
used statistical tool designed to detect periodic signals in unevenly spaced
observations. The base LombScargle provides an
interface for several implementations of the Lomb-Scargle periodogram. However,
LombScargle only handles a single band of data.
The LombScargleMultiband class adapts this
interface to handle multiband data (where multiple bands/filters are present).
The code here is adapted from the astroml package ([3], [4]) and the
gatspy package ([5], [6]), but conforms closely to the design paradigms
established in LombScargle. For a detailed
practical discussion of the Multiband Lomb-Scargle periodogram, which guided
the development of this class, see
Periodograms for Multiband Astronomical Time Series [6].
Basic Usage#
Note
As in LombScargle, frequencies in
LombScargleMultiband are not
angular frequencies, but rather frequencies of oscillation (i.e., number of
cycles per unit time).
The Lomb-Scargle Multiband periodogram is designed to detect periodic signals in unevenly spaced observations with multiple bands of data present.
Example#
To detect periodic signals in unevenly spaced observations, consider the following multiband data, where 5 bands (u, g, r, i, and z) have 60 datapoints each.
>>> import numpy as np
>>> t = []
>>> y = []
>>> bands = []
>>> dy = []
>>> N=60
>>> for i, band in enumerate(['u','g','r','i','z']):
... rng = np.random.default_rng(i)
... t_band = 300 * rng.random(N)
... y_band = 3 + 2 * np.sin(2 * np.pi * t_band)
... dy_band = 0.01 * (0.5 + rng.random(N)) # uncertainties
... y_band += dy_band * rng.standard_normal(N)
... t += list(t_band)
... y += list(y_band)
... dy += list(dy_band)
... bands += [band] * N
The Lomb-Scargle periodogram, evaluated at frequencies chosen
automatically based on the input data, can be computed as follows
using the LombScargleMultiband class,
with the bands argument being the sole difference in comparison
to the LombScargle interface:
>>> from astropy.timeseries import LombScargleMultiband
>>> frequency,power = LombScargleMultiband(t, y, bands, dy).autopower()
Plotting the result with Matplotlib gives:
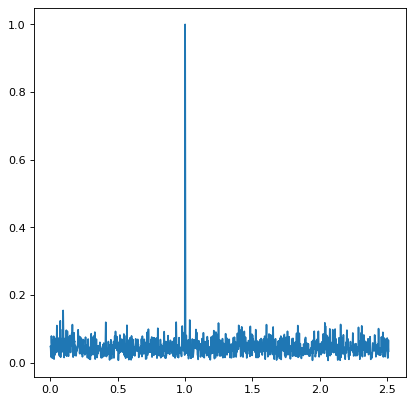
The periodogram shows a clear spike at a frequency of 1 cycle per unit time,
as we would expect from the data we constructed. The resulting power is a
single array, with combined input from each of the bands dependent upon the
implementation chosen in the method keyword.
Periodograms from TimeSeries objects#
LombScargleMultiband is able to operate on
TimeSeries objects, provided the
TimeSeries object meets a formatting requirement.
The requirement is that the flux (or magnitudes) and errors for each band are
provided in separate columns. If instead, your
TimeSeries object has a singular flux column with
an associated band label column, these columns may be passed directly to
LombScargleMultiband as 1-d arrays.
Example#
Consider the following generator code for a
TimeSeries object where timeseries data is
populated for three photometric bands (g,r,i).
>>> from astropy.timeseries import LombScargleMultiband, TimeSeries
>>> from astropy.table import MaskedColumn
>>> import numpy as np
>>> import astropy.units as u
>>> rng = np.random.default_rng(1)
>>> deltas = 240 * rng.random(180)
>>> ts1 = TimeSeries(time_start="2011-01-01T00:00:00",
... time_delta=deltas*u.minute)
>>> # g band fluxes
>>> g_flux = [0] * 180 * u.mJy
>>> g_err = [0] * 180 * u.mJy
>>> y_g = np.round(3 + 2 * np.sin(10 * np.pi * ts1['time'].mjd[0:60]),3)
>>> dy_g = np.round(0.01 * (0.5 + rng.random(60)), 3) # uncertainties
>>> g_flux.value[0:60] = y_g
>>> g_err.value[0:60] = dy_g
>>> ts1["g_flux"] = MaskedColumn(g_flux, mask=[False]*60+[True]*120)
>>> ts1["g_err"] = MaskedColumn(g_err, mask=[False]*60+[True]*120)
>>> # r band fluxes
>>> r_flux = [0] * 180 * u.mJy
>>> r_err = [0] * 180 * u.mJy
>>> y_r = np.round(3 + 2 * np.sin(10 * np.pi * ts1['time'].mjd[60:120]),3)
>>> dy_r = np.round(0.01 * (0.5 + rng.random(60)), 3) # uncertainties
>>> r_flux.value[60:120] = y_r
>>> r_err.value[60:120] = dy_r
>>> ts1['r_flux'] = MaskedColumn(r_flux, mask=[True]*60+[False]*60+[True]*60)
>>> ts1['r_err'] = MaskedColumn(r_err, mask=[True]*60+[False]*60+[True]*60)
>>> # i band fluxes
>>> i_flux = [0] * 180 * u.mJy
>>> i_err = [0] * 180 * u.mJy
>>> y_i = np.round(3 + 2 * np.sin(10 * np.pi * ts1['time'].mjd[120:]),3)
>>> dy_i = np.round(0.01 * (0.5 + rng.random(60)), 3) # uncertainties
>>> i_flux.value[120:] = y_i
>>> i_err.value[120:] = dy_i
>>> ts1["i_flux"] = MaskedColumn(i_flux, mask=[True]*120+[False]*60)
>>> ts1["i_err"] = MaskedColumn(i_err, mask=[True]*120+[False]*60)
>>> ts1
<TimeSeries length=180>
time g_flux g_err r_flux r_err i_flux i_err
mJy mJy mJy mJy mJy mJy
Time float64 float64 float64 float64 float64 float64
----------------------- ------- ------- ------- ------- ------- -------
2011-01-01T00:00:00.000 3.0 0.012 ——— ——— ——— ———
2011-01-01T02:02:50.231 3.891 0.009 ——— ——— ——— ———
2011-01-01T05:50:56.909 4.961 0.007 ——— ——— ——— ———
2011-01-01T06:25:32.807 4.697 0.014 ——— ——— ——— ———
2011-01-01T10:13:13.359 4.451 0.005 ——— ——— ——— ———
2011-01-01T11:28:03.732 4.283 0.008 ——— ——— ——— ———
2011-01-01T13:09:39.633 1.003 0.015 ——— ——— ——— ———
2011-01-01T16:28:18.550 3.833 0.008 ——— ——— ——— ———
2011-01-01T18:06:31.018 1.02 0.013 ——— ——— ——— ———
... ... ... ... ... ... ...
2011-01-15T16:03:17.207 ——— ——— ——— ——— 4.656 0.014
2011-01-15T17:29:38.139 ——— ——— ——— ——— 1.423 0.01
2011-01-15T20:03:35.935 ——— ——— ——— ——— 4.805 0.008
2011-01-15T21:35:02.069 ——— ——— ——— ——— 3.042 0.007
2011-01-15T23:06:35.567 ——— ——— ——— ——— 1.162 0.01
2011-01-16T01:07:30.330 ——— ——— ——— ——— 4.99 0.009
2011-01-16T01:11:31.138 ——— ——— ——— ——— 5.0 0.011
2011-01-16T03:09:58.569 ——— ——— ——— ——— 1.314 0.01
2011-01-16T07:03:09.586 ——— ——— ——— ——— 3.383 0.005
Our timeseries data is set up to be asynchronous, where a given timestamp
corresponds to a measurement in a single band. However, if your data instead
has one timestamp per multiple band measurements, or a mixture,
LombScargleMultiband will still be able to
operate on it.
To operate on the example TimeSeries,
LombScargleMultiband has a loader function, as
follows:
>>> ls = LombScargleMultiband.from_timeseries(ts1, signal_column=['g_flux', 'r_flux', 'i_flux'],
... uncertainty_column=['g_err', 'r_err', 'i_err'],
... band_labels=['g', 'r', 'i'])
signal_column requires a list of columns that correspond to the flux
or magnitude measurements in each band. uncertainty_column and
band_labels are optional, but if specified must be lists of equal size to
signal_column. uncertainty_column specifies the columns containing the
associated errors per band, while band_labels provides the labels to use
for each photometric band. From here,
LombScargleMultiband can be worked with as normal.
For example:
>>> frequency,power = ls.autopower()
Consistencies with LombScargle#
LombScargleMultiband is an inherited class of
LombScargle, and was developed to provide as
similar of an interface to LombScargle as
possible. From this, there are several core aspects of
LombScargle that remain true for
LombScargleMultiband.
Measurement Uncertainties#
The LombScargleMultiband interface can also handle
data with measurement uncertainties. As shown in the example above.
Periodograms and Units#
The LombScargleMultiband interface properly
handles Quantity objects with units attached,
and will validate the inputs to make sure units are appropriate.
Specifying the Frequency Grid#
As shown above, the autopower()
method automatically determines a frequency grid, using
autofrequency(). The tunable parameters
are identical to those shown for LombScargle. And
likewise, a custom frequency grid may be supplied directly to the
power() function.
Example#
>>> frequency = np.linspace(0, 2, 1000)
>>> power = LombScargleMultiband(t, y, bands, dy).power(frequency)
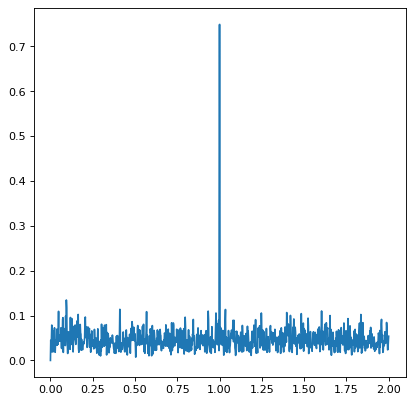
Periodogram Implementations#
Two implementations of the Multiband Lomb-Scargle Periodogram are available
within LombScargleMultiband, flexible and
fast, which are selectable via the
power() method’s
method parameter. flexible is a direct port of the LombScargleMultiband
algorithm used in the gatspy gatspy package. It constructs a common model,
and an offset model per individual band. It then applies regularization to the
resulting model to constrain complexity, resulting in a flexible model for any
given multiband timeseries dataset. As it’s name implies, fast is
potentially quicker alternative that fits each band independently and combines
them by weight. The independent band-by-band fits leverage
LombScargle. As a result the sb_method
parameter is available in
power() to choose the
single-band method used in power() for
each band. Keep in mind that the speed of fast is dependent on the
underlying speed of the choice of sb_method.
Example#
flexible:
>>> frequency, power = LombScargleMultiband(t,y,bands,dy).autopower(method='flexible')
fast, with fast also chosen as the
power() method:
>>> frequency, power = LombScargleMultiband(t,y,bands,dy).autopower(method='fast', sb_method='fast')
The Multiband Lomb-Scargle Model#
The model() method fits a
sinusoidal model to the data at a chosen frequency. The sinusoidal model
complexity is tunable via the nterms_base and nterms_band parameters.
These control the number of sinusoidal terms available to the base model
(common to all bands) and the number of sinusoidal terms available to each
bands offset model.
Note
Either of nterms_base and nterms_band may be set to 0, though not
both. The case when nterms_base =0 and nterms_band =1 is a special
case referred to as the multi-phase model, where the base model is reduced
to a simple offset, and therefore the bands are solved independently (a
single-band fit). Further discussed in
Periodograms for Multiband Astronomical Time Series [6]
Example#
The following example uses the same data as above.
autopower() is used to return
the periodogram, and we can select the frequency at which the power is maximum
for our model:
>>> model = LombScargleMultiband(t, y, bands, dy, nterms_base=1, nterms_band=1)
>>> frequency, power = model.autopower(method='flexible')
>>> freq_maxpower = frequency[np.argmax(power)]
We can then model based on the found frequency, and time (phased by the frequency):
>>> t_phase = np.linspace(0, 1/freq_maxpower, 100)
>>> y_fit = model.model(t_phase, freq_maxpower)
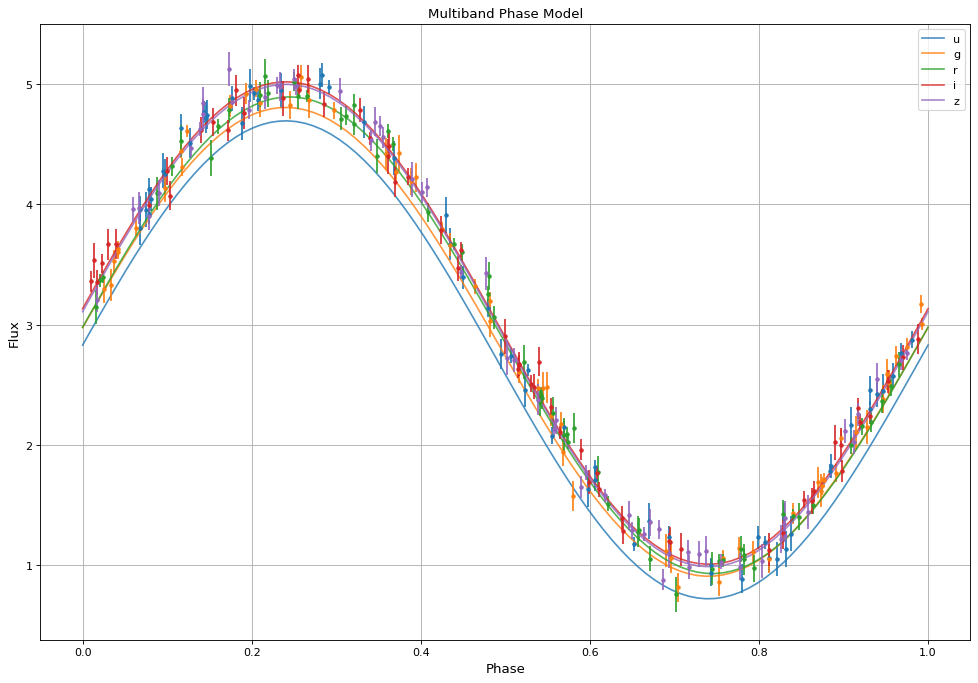
The resulting fit is then of shape (number of bands, number of timesteps), or (5,100) in this particular case. By plotting the result, we see the model has recovered the expected sinusoid recovered at the correct frequency:
False Alarm Probabilities#
Unlike LombScargle,
LombScargleMultiband does not have False Alarm
Probabilities implemented. The algorithms available for
LombScargle are valid only for single term
periodograms, which is rarely valid for models in the Multiband case.
