N-Dimensional Datasets (astropy.nddata)#
Introduction#
The nddata package provides classes to represent images and other
gridded data, some essential functions for manipulating images, and the
infrastructure for package developers who wish to include support for the
image classes. This subpackage was developed based on APE 7.
Getting Started#
NDData#
The primary purpose of NDData is to act as a container for
data, metadata, and other related information like a mask.
An NDData object can be instantiated by passing it an
n-dimensional numpy array:
>>> import numpy as np
>>> from astropy.nddata import NDData
>>> array = np.zeros((12, 12, 12)) # a 3-dimensional array with all zeros
>>> ndd1 = NDData(array)
Or something that can be converted to a numpy.ndarray:
>>> ndd2 = NDData([1, 2, 3, 4])
>>> ndd2
NDData([1, 2, 3, 4])
And can be accessed again via the data attribute:
>>> ndd2.data
array([1, 2, 3, 4])
It also supports additional properties like a unit or mask for the
data, a wcs (World Coordinate System) and uncertainty of the data and
additional meta attributes:
>>> data = np.array([1,2,3,4])
>>> mask = data > 2
>>> unit = 'erg / s'
>>> from astropy.nddata import StdDevUncertainty
>>> uncertainty = StdDevUncertainty(np.sqrt(data)) # representing standard deviation
>>> meta = {'object': 'fictional data.'}
>>> ndd = NDData(data, mask=mask, unit=unit, uncertainty=uncertainty,
... meta=meta)
>>> ndd
NDData([1, 2, —, —], unit='erg / s')
The representation only displays the data; the other attributes need to be
accessed directly, for example, ndd.mask to access the mask.
NDDataRef#
Building upon this pure container, NDDataRef implements:
A
readandwritemethod to accessastropy’s unified file I/O interface.Simple arithmetic like addition, subtraction, division, and multiplication.
Slicing.
Instances are created in the same way:
>>> from astropy.nddata import NDDataRef
>>> ndd = NDDataRef(ndd)
>>> ndd
NDDataRef([1, 2, —, —], unit='erg / s')
But also support arithmetic (NDData Arithmetic) like addition:
>>> import astropy.units as u
>>> ndd2 = ndd.add([4, -3.5, 3, 2.5] * u.erg / u.s)
>>> ndd2
NDDataRef([ 5. , -1.5, ———, ———], unit='erg / s')
Because these operations have a wide range of options, these are not available
using arithmetic operators like +.
Slicing or indexing (Slicing and Indexing NDData) is possible (with warnings issued if some attribute cannot be sliced):
>>> ndd2[2:] # discard the first two elements
NDDataRef([———, ———], unit='erg / s')
>>> ndd2[1] # get the second element
NDDataRef(-1.5, unit='erg / s')
Working with Two-Dimensional Data Like Images#
Though the nddata package supports any kind of gridded data, this
introduction will focus on the use of nddata for two-dimensional
images. To get started, we will construct a two-dimensional image with a few
sources, some Gaussian noise, and a “cosmic ray” which we will later mask out.
Examples#
First, construct a two-dimensional image with a few sources, some Gaussian noise, and a “cosmic ray”:
>>> import numpy as np
>>> from astropy.modeling.models import Gaussian2D
>>> rng = np.random.default_rng()
>>> y, x = np.mgrid[0:500, 0:600]
>>> data = (Gaussian2D(1, 150, 100, 20, 10, theta=0.5)(x, y) +
... Gaussian2D(0.5, 400, 300, 8, 12, theta=1.2)(x,y) +
... Gaussian2D(0.75, 250, 400, 5, 7, theta=0.23)(x,y) +
... Gaussian2D(0.9, 525, 150, 3, 3)(x,y) +
... Gaussian2D(0.6, 200, 225, 3, 3)(x,y))
>>> data += 0.01 * rng.standard_normal((500, 600))
>>> cosmic_ray_value = 0.997
>>> data[100, 300:310] = cosmic_ray_value
This image has a large “galaxy” in the lower left and the “cosmic ray” is the horizontal line in the lower middle of the image:
>>> import matplotlib.pyplot as plt
>>> plt.imshow(data, origin='lower')

The “cosmic ray” can be masked out in this test image, like this:
>>> mask = (data == cosmic_ray_value)
CCDData Class for Images#
The CCDData object, like the other objects in this package,
can store the data, a mask, and metadata. The CCDData object
requires that a unit be specified:
>>> from astropy.nddata import CCDData
>>> ccd = CCDData(data, mask=mask,
... meta={'object': 'fake galaxy', 'filter': 'R'},
... unit='adu')
Slicing#
Slicing works the way you would expect with the mask and, if present, WCS, sliced appropriately:
>>> ccd2 = ccd[:200, :]
>>> ccd2.data.shape
(200, 600)
>>> ccd2.mask.shape
(200, 600)
>>> # Show the mask in a region around the cosmic ray:
>>> ccd2.mask[99:102, 299:311]
array([[False, False, False, False, False, False, False, False, False,
False, False, False],
[False, True, True, True, True, True, True, True, True,
True, True, False],
[False, False, False, False, False, False, False, False, False,
False, False, False]]...)
For many applications it may be more convenient to use
Cutout2D, described in image_utilities.
Image Arithmetic, Including Uncertainty#
Methods are provided for basic arithmetic operations between images, including
propagation of uncertainties. Three uncertainty types are supported: variance
(VarianceUncertainty), standard deviation
(StdDevUncertainty), and inverse variance
(InverseVariance).
Examples#
This example creates an uncertainty that is Poisson error, stored as a variance:
>>> from astropy.nddata import VarianceUncertainty
>>> poisson_noise = np.ma.sqrt(np.ma.abs(ccd.data))
>>> ccd.uncertainty = VarianceUncertainty(poisson_noise ** 2)
As a convenience, the uncertainty can also be set with a numpy array. In
that case, the uncertainty is assumed to be the standard deviation:
>>> ccd.uncertainty = poisson_noise
INFO: array provided for uncertainty; assuming it is a StdDevUncertainty. [astropy.nddata.ccddata]
If we make a copy of the image and add that to the original, the uncertainty changes as expected:
>>> ccd2 = ccd.copy()
>>> added_ccds = ccd.add(ccd2, handle_meta='first_found')
>>> added_ccds.uncertainty.array[0, 0] / ccd.uncertainty.array[0, 0] / np.sqrt(2)
0.99999999999999989
Reading and Writing#
A CCDData can be saved to a FITS file:
>>> ccd.write('test_file.fits')
And can also be read in from a FITS file:
>>> ccd2 = CCDData.read('test_file.fits')
Note the unit is stored in the BUNIT keyword in the header on saving, and is
read from the header if it is present.
Detailed help on the available keyword arguments for reading and writing
can be obtained via the help() method as follows:
>>> CCDData.read.help('fits') # Get help on the CCDData FITS reader
>>> CCDData.writer.help('fits') # Get help on the CCDData FITS writer
Image Utilities#
Cutouts#
Though slicing directly is one way to extract a subframe,
Cutout2D provides more convenient access to cutouts from the
data.
Examples#
This example pulls out the large “galaxy” in the lower left of the image, with
the center of the cutout at position:
>>> from astropy.nddata import Cutout2D
>>> position = (149.7, 100.1)
>>> size = (81, 101) # pixels
>>> cutout = Cutout2D(ccd, position, size)
>>> plt.imshow(cutout.data, origin='lower')
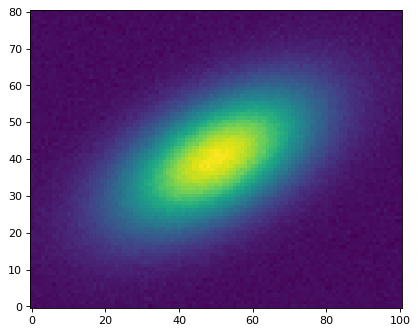
This cutout can also plot itself on the original image:
>>> plt.imshow(ccd, origin='lower')
>>> cutout.plot_on_original(color='white')

The cutout also provides methods for finding pixel coordinates in the original
or in the cutout; recall that position is the center of the cutout in the
original image:
>>> position
(149.7, 100.1)
>>> cutout.to_cutout_position(position)
(49.7, 40.099999999999994)
>>> cutout.to_original_position((49.7, 40.099999999999994))
(149.7, 100.1)
For more details, including constructing a cutout from World Coordinates and the options for handling cutouts that go beyond the bounds of the original image, see 2D Cutout Images.
Image Resizing#
The functions block_reduce and
block_replicate resize images.
Example#
This example reduces the size of the image by a factor of 4. Note that the
result is a numpy.ndarray; the mask, metadata, etc. are discarded:
>>> from astropy.nddata import block_reduce, block_replicate
>>> smaller = block_reduce(ccd, 4)
>>> smaller
array(...)
>>> plt.imshow(smaller, origin='lower')

By default, both block_reduce and
block_replicate conserve flux.
Other Image Classes#
There are two less restrictive classes, NDDataArray and
NDDataRef, that can be used to hold image data. They are
primarily of interest to those who may want to create their own image class by
subclassing from one of the classes in the nddata package. The main
differences between them are:
NDDataRefcan be sliced and has methods for basic arithmetic operations, but the user needs to use one of the uncertainty classes to define an uncertainty. See NDDataRef for more detail. Most of its properties must be set when the object is created because they are not mutable.NDDataArrayextendsNDDataRefby adding the methods necessary for it to behave like anumpyarray in expressions and adds setters for several properties. It lacks the ability to automatically recognize and read data from FITS files and does not attempt to automatically set the WCS property.CCDDataextendsNDDataArrayby setting up a default uncertainty class, setting up straightforward read/write to FITS files, and automatically setting up a WCS property.
More General Gridded Data Classes#
There are two generic classes in the nddata package that are of
interest primarily to users who either need a custom image class that goes
beyond the classes discussed so far, or who are working with gridded data that
is not an image.
NDDatais a container class for holding general gridded data. It includes a handful of basic attributes, but no slicing or arithmetic. More information about this class is in NDData.NDDataBaseis an abstract base class that developers of new gridded data classes can subclass to declare that the new class follows theNDDatainterface. More details are in Subclassing.
Additional Examples#
The list of packages below that use the nddata framework is intended to be
useful to either users writing their own image classes or those looking
for an image class that goes beyond what CCDData does.
The SunPy project uses
NDDataas the foundation for its Map classes.The class
NDDataRefis used in specutils as the basis for Spectrum1D, which adds several methods useful for spectra.The package ndmapper, which makes it easy to build reduction pipelines for optical data, uses
NDDataArrayas its image object.The package ccdproc uses the
CCDDataclass throughout for implementing optical/IR image reduction.
Using nddata#
Performance Tips#
Using the uncertainty class
VarianceUncertaintywill be somewhat more efficient than the other two uncertainty classes,InverseVarianceandStdDevUncertainty. The latter two are converted to variance for the purposes of error propagation and then converted from variance back to the original uncertainty type. The performance difference should be small.When possible, mask values by setting them to
np.nanand use thenumpyfunctions and methods that automatically excludenp.nan, likenp.nanmedianandnp.nanstd. This will typically be much faster than usingnumpy.ma.MaskedArray.
